Innovations Track
Quantifying the Impact of Dataset Shifts in Deep Learning-based Dynamic MRI Reconstruction: Need for Spatially Localized Performance Metrics

M. Berk Sahin
PhD student
Purdue University
West Lafayette, Indiana, United States
M. Berk Sahin
PhD student
Purdue University
West Lafayette, Indiana, United States- DY
Dilek M. Yalcinkaya, MSc
PhD Candidate
Purdue University
West Lafayette, Indiana, United States 
Hazar Benan Unal, PhD
Postdoctoral Research Associate
Laboratory for Translational Imaging of Microcirculation, Purdue University
Indianapolis, Indiana, United States
Khalid Youssef, PhD, MSc
Assistant Professor
Indiana University, Department of Radiology and Imaging Sciences
Indianapolis, Indiana, United States- AH
Abolfazl Hashemi, PhD, MSc, BSc
Assistant Professor
Purdue University, Indiana, United States - BS
Behzad Sharif, PhD
Associate Professor
Purdue University
Indianapolis, Indiana, United States
Presenting Author(s)
Primary Author(s)
Co-Author(s)
Methods: We trained two state-of-the-art DL reconstruction methods, a 2D-static VarNet [2] and a 2D+time VarNet [3], on cardiac perfusion (n = 142) and cine (n = 146) data separately. We used retrospective subsampling for training and evaluation on multiple test sets representing different dataset shifts. Reconstruction performance was quantified using global and localized NRMSE. The overall workflow is outlined in Fig. 1A. Localized NRMSE was calculated by applying an already trained DL-based localizer [4] to identify the cardiac region and calculating NRMSE only within this area over time. Global and localized NRMSE were then compared across various dataset shifts summarized in Fig. 1B.
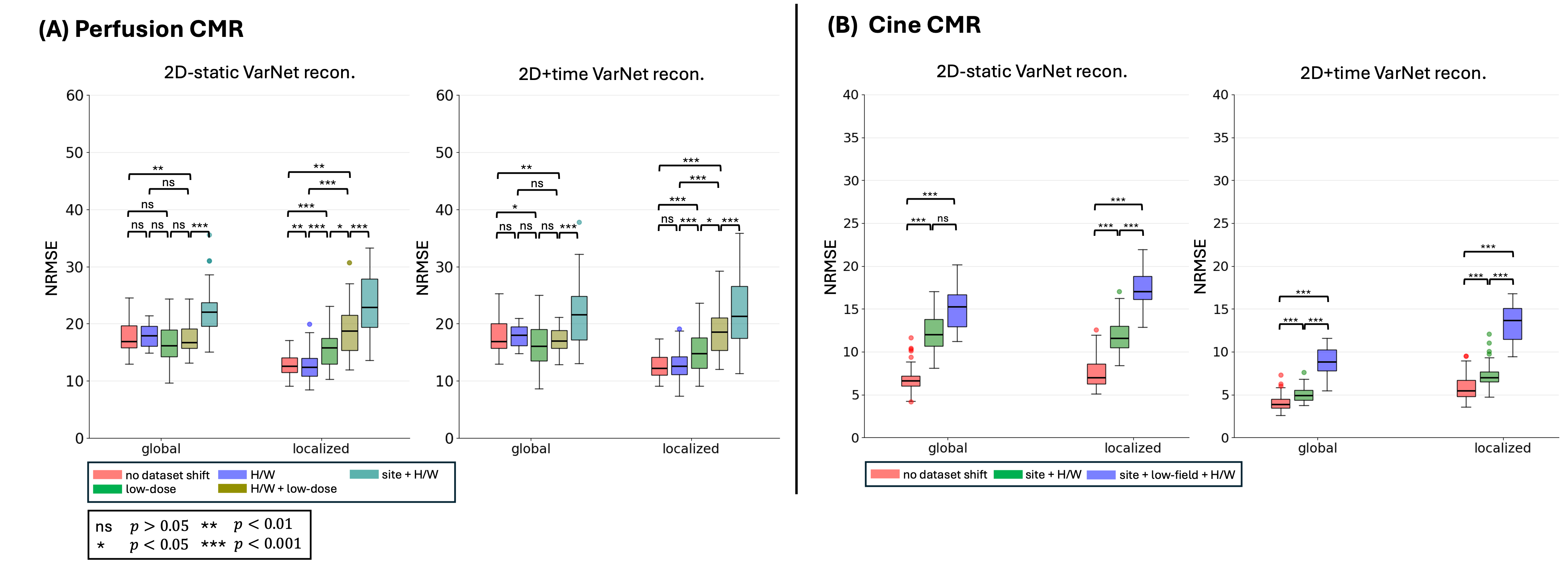
Results: In perfusion reconstructions of the same patient in Fig. 2A, localized NRMSE captured much larger error differences between the full-dose (no dataset shift) and low-dose (dataset shift) reconstructions than global NRMSE, both in the x-y view (Fig. 2A) and in the y-t cut (Fig. 2B). For example, in 2D-static reconstructions under the low-dose protocol, global NRMSE increased by only 0.4% in the x–y view and 0.3% in the y–t cut, whereas localized NRMSE increased by 4.7% and 3.4%, respectively. This pattern is consistent with the cumulative results in Fig. 3. Global NRMSE showed no statistically significant variation across test sets (p >0.05). In contrast, localized NRMSE consistently separated (p< 0.05) reconstruction errors across dataset shifts in both perfusion and cine CMR.
Conclusion: Our proposed localized NRMSE metric provides a more sensitive and clinically relevant assessment of dataset shift–related degradations in DL-based reconstruction. Unlike global NRMSE, which remain largely insensitive to dataset shifts, localized NRMSE consistently detected significant differences across shifts. This improved sensitivity can help guide the design of reconstruction methods that generalize more robustly to unseen data, ultimately enhancing their reliability and clinical applicability.
Figure 1: (A) Overview of the proposed pipeline for evaluating different dataset shift types using global and localized reconstruction error. (B) Description of the cardiac perfusion and cine CMR datasets used in this study. In panel (A), training and test stages of the pipeline are shown on myocardial perfusion images (top and bottom, respectively), with identical steps applied to the cine dataset. Training Stage – Step 1: The internal data were reconstructed using time-interleaved GRAPPA [5] and served as reference image. Training Stage – Step 2 and 3: Reconstructed data were divided into training, validation and internal test (i.e., “no dataset shift”) sets. The training and validation sets were used for hyperparameter tuning of two DL-based reconstruction models (2D-static [2] and 2D+time VarNet [3]). Both were trained on retrospectively subsampled k-space from the reference images at effective acceleration rate of 3.5 using Cartesian sampling with a 10% ACS region. For each DL method, the version that performed the best over validation set were used for final testing. Test Stage – Step 4: Following the same data preparation as in Step 1, raw k-spaces from different test sets, introducing dataset shifts in scanner hardware, contrast dose (full vs. low), and imaging site, were retrospectively subsampled at effective acceleration rate of 3.5 and the subsampled k-spaces were reconstructed with the trained DL-based models. Test Stage – Step 5: A pre-trained DL-based localizer [4] was used to extract the cardiac region, indicated by red rectangle, in reconstructed series. Test Stage – Step 6: Global and localized NRMSE values were calculated over the entire reconstructed series (green) and the cardiac region (red). As summarized in panel (B), various dataset shifts across both modalities were tested to examine their impact on localized and global NRMSE. For perfusion, we employed in-house datasets acquired across multiple sites and scanners (H/W), including both full-dose and low-dose contrast protocols. For cine, we utilized publicly available data from CMRxRecon [6] and OCMR [7] with site and scanner variations.
.png) Figure 2: (A) Myocardial perfusion representative reconstruction results from two different scans of the same patient under no dataset shift and dataset shift due to a low-dose contrast protocol, highlighting the importance of using localized error metric to accurately assess the impact of dataset shift. From left to right, the first column shows the artifact-free scanner reconstruction, which is considered reference image. Subsequent columns display reconstructions from retrospectively subsampled data at effective acceleration rate of 3.5 using a Cartesian sampling with 10% calibration lines. These include a zero-filled reconstruction (inverse FFT with missing data set to zero), GRAPPA [5], and two DL-based methods (2D-static and 2D+time VarNet). Global (green) and localized (red) NRMSE values are shown at the top-left and top-right of each reconstruction. Under the low-dose protocol, global NRMSE increased by 0.4% for 2D-static and 0.1% for 2D+time reconstructions, while localized NRMSE increased by 4.7% and 3.6%, respectively. (B) Myocardial perfusion representative y-t cuts corresponding to the scans in (A). The first column shows the x-y view of the reference image with the cardiac region (red box) generated by a pre-trained localizer. Green dashed lines show the y-t cut location whereas red dashed lines follow the localization box on y-t cut over time. Global NRMSE was calculated over the full y–t cut, whereas localized NRMSE was calculated only within the segment of the y–t cut that falls inside the red box. Under the low-dose protocol dataset shift, global (green, top-left) NRMSE changed by 0.3% for 2D-static and 0.7% for 2D+time reconstructions, while localized NRMSE changed by 3.4% and 3.1%, respectively.
Figure 2: (A) Myocardial perfusion representative reconstruction results from two different scans of the same patient under no dataset shift and dataset shift due to a low-dose contrast protocol, highlighting the importance of using localized error metric to accurately assess the impact of dataset shift. From left to right, the first column shows the artifact-free scanner reconstruction, which is considered reference image. Subsequent columns display reconstructions from retrospectively subsampled data at effective acceleration rate of 3.5 using a Cartesian sampling with 10% calibration lines. These include a zero-filled reconstruction (inverse FFT with missing data set to zero), GRAPPA [5], and two DL-based methods (2D-static and 2D+time VarNet). Global (green) and localized (red) NRMSE values are shown at the top-left and top-right of each reconstruction. Under the low-dose protocol, global NRMSE increased by 0.4% for 2D-static and 0.1% for 2D+time reconstructions, while localized NRMSE increased by 4.7% and 3.6%, respectively. (B) Myocardial perfusion representative y-t cuts corresponding to the scans in (A). The first column shows the x-y view of the reference image with the cardiac region (red box) generated by a pre-trained localizer. Green dashed lines show the y-t cut location whereas red dashed lines follow the localization box on y-t cut over time. Global NRMSE was calculated over the full y–t cut, whereas localized NRMSE was calculated only within the segment of the y–t cut that falls inside the red box. Under the low-dose protocol dataset shift, global (green, top-left) NRMSE changed by 0.3% for 2D-static and 0.7% for 2D+time reconstructions, while localized NRMSE changed by 3.4% and 3.1%, respectively.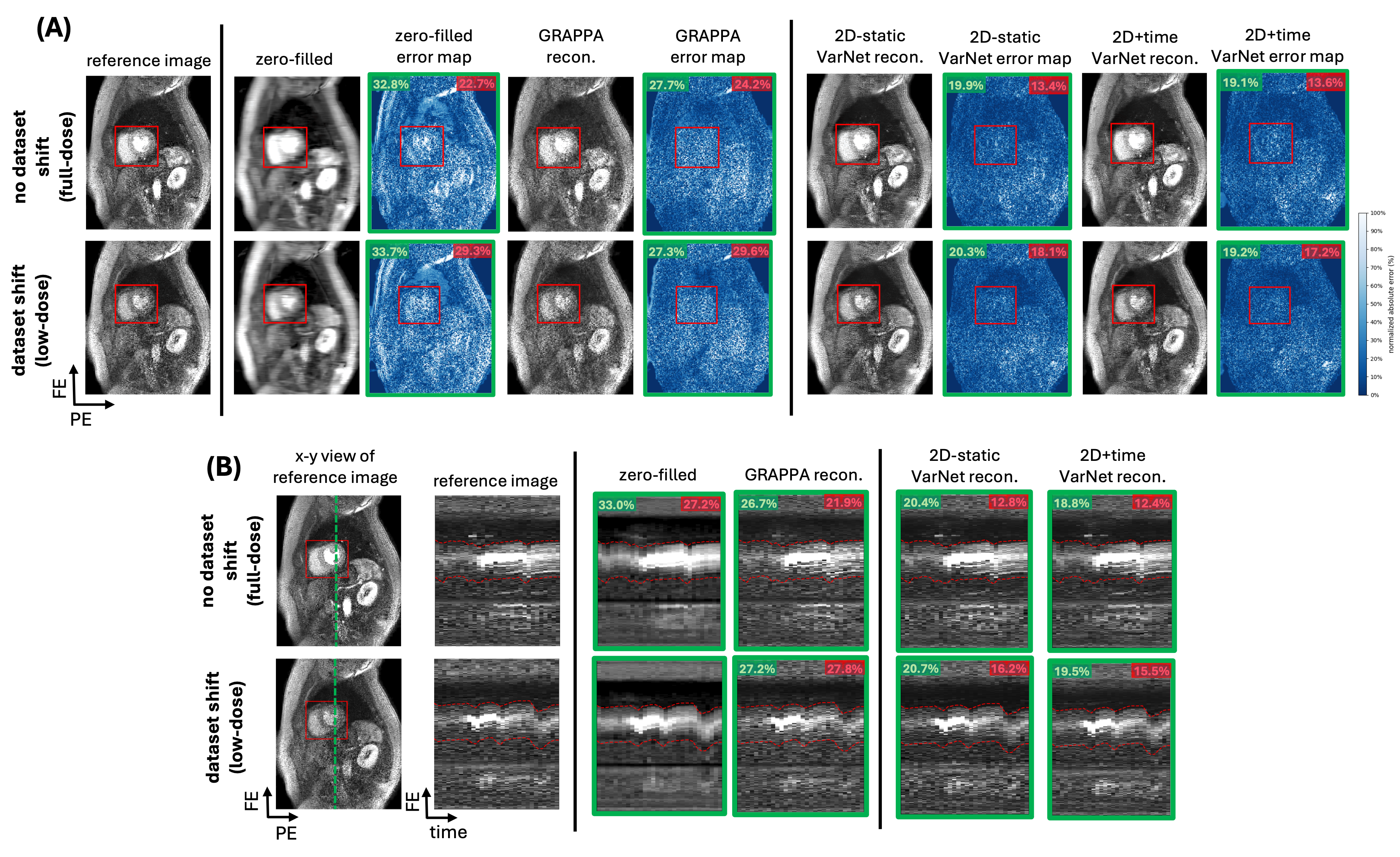 Figure 3: DL-based cardiac perfusion and cine reconstructions under different dataset shifts, illustrating that localized errors capture model generalization more effectively than global errors. Panels (A) and (B) show cumulative reconstruction results for perfusion and cine CMR at effective acceleration of 3.5 retrospectively, for two DL-based methods (2D-static and 2D+time VarNet), trained separately and evaluated on different test sets including “no dataset shift” case from Fig. 2B. Global NRMSE, computed over the entire image, shows little variation across test sets and statistically not significant (p>0.05). In contrast, localized NRMSE, computed within the cardiac region from the DL-based localizer, consistently separates the reconstruction error in different test sets and yields more statistically significant differences. This pattern is observed consistently across both DL models and both imaging modalities, demonstrating that localized NRMSE is more sensitive to dataset shifts than global NRMSE.
Figure 3: DL-based cardiac perfusion and cine reconstructions under different dataset shifts, illustrating that localized errors capture model generalization more effectively than global errors. Panels (A) and (B) show cumulative reconstruction results for perfusion and cine CMR at effective acceleration of 3.5 retrospectively, for two DL-based methods (2D-static and 2D+time VarNet), trained separately and evaluated on different test sets including “no dataset shift” case from Fig. 2B. Global NRMSE, computed over the entire image, shows little variation across test sets and statistically not significant (p>0.05). In contrast, localized NRMSE, computed within the cardiac region from the DL-based localizer, consistently separates the reconstruction error in different test sets and yields more statistically significant differences. This pattern is observed consistently across both DL models and both imaging modalities, demonstrating that localized NRMSE is more sensitive to dataset shifts than global NRMSE.